Introduction to Machine Learning
CMSC 173 - Module 00
Noel Jeffrey Pinton
Department of Computer Science
University of the Philippines Cebu
Course Overview
Topics Covered
- What is Machine Learning?
- Types of Learning: Supervised, Unsupervised, Reinforcement
- The ML Pipeline
- Bias-Variance Tradeoff
- Best Practices & Ethics
What is Machine Learning?
Formal Definition
Machine Learning (ML) is the science of getting computers to learn and act like humans do, and improve their learning over time in autonomous fashion, by feeding them data and information.Key Characteristics
- Learning from data without explicit programming
- Improving performance with experience
- Discovering patterns in complex datasets
- Making predictions or decisions
Traditional Programming vs ML
Traditional:\\ Rules + Data → Answers Machine Learning:\\ Data + Answers → RulesCore Insight
ML finds the rules automatically from examples!Traditional Programming vs ML
| Aspect | Traditional | Machine Learning |
|---|---|---|
| Input | Rules + Data | Data + Labels |
| Output | Answers | Rules/Model |
| Example | if price > 1000: expensive | Learn threshold from examples |
Historical Context
"A computer would deserve to be called intelligent if it could deceive a human into believing that it was human." --- Alan TuringMajor Milestones
- 1950s: Alan Turing - "Can machines think?"
- 1957: Perceptron (Frank Rosenblatt)
- 1986: Backpropagation popularized
- 1990s: Support Vector Machines
- 1997: Deep Blue defeats Kasparov
- 2006: Deep Learning renaissance
- 2012: AlexNet wins ImageNet
- 2016: AlphaGo defeats Lee Sedol
- 2020s: Large Language Models
The Three AI Winters
Periods of reduced funding and interest:- 1970s: Perceptron limitations
- 1987-1993: Expert systems fail
- Post-2000: AI hype deflation
Current Era
We're in the Deep Learning Revolution:- Big data availability
- GPU acceleration
- Novel architectures (Transformers)
- Widespread deployment
Real-World Applications
"Machine learning is the last invention that humanity will ever need to make." --- Nick BostromComputer Vision
- Medical image diagnosis
- Autonomous vehicles
- Facial recognition
- Object detection & tracking
- Image generation (DALL-E, Midjourney)
Natural Language Processing
- Machine translation
- Chatbots & virtual assistants
- Sentiment analysis
- Text summarization
- Question answering
Other Domains
- Finance: Fraud detection, trading
- Healthcare: Drug discovery, medicine
- E-commerce: Recommendations
- Gaming: AI opponents
- Manufacturing: Quality control
- Agriculture: Crop monitoring
Impact
ML is transforming every industry!Learning Objectives
By the end of this course, you will be able to:
- Understand the fundamental concepts and mathematical foundations of machine learning
- Distinguish between different types of learning paradigms (supervised, unsupervised, etc.)
- Implement core ML algorithms from scratch using Python
- Apply appropriate ML techniques to real-world problems
- Evaluate model performance using rigorous metrics
- Analyze the theoretical properties of learning algorithms
- Compare different approaches and select optimal methods
- Understand state-of-the-art techniques in deep learning
Prerequisites
CMSC 170: Linear algebra, probability theory, calculus, Python programmingMachine Learning Taxonomy
See visual diagram in lecture materials
Supervised Learning
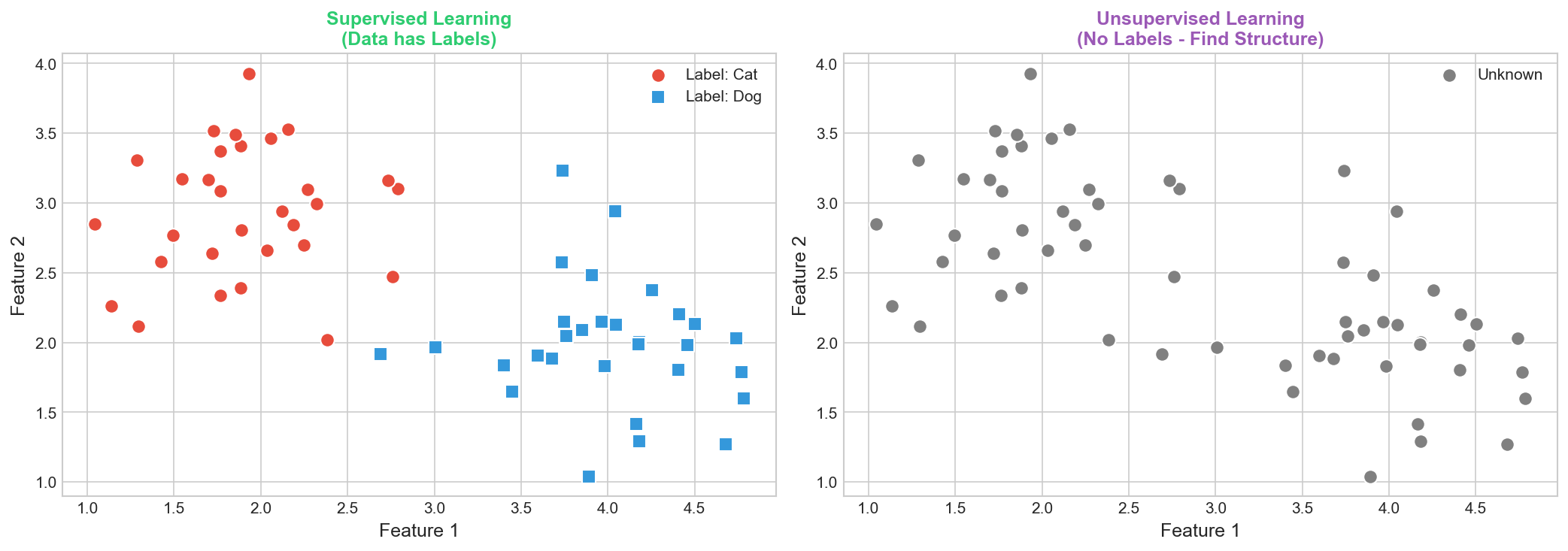
Definition: Learning from labeled data
- Input: $\mathbf{x} \in \mathbb{R}^d$
- Output: Label $y$
- Goal: Learn $f(\mathbf{x}) \approx y$
Two Main Tasks
- Regression: $y \in \mathbb{R}$
- Classification: $y \in \{1,...,K\}$
Supervised Learning: Training
Training Process
Given $\mathcal{D} = \{(\mathbf{x}_i, y_i)\}_{i=1}^n$:
- Choose hypothesis class $\mathcal{H}$
- Define loss $\mathcal{L}(y, \hat{y})$
- Minimize empirical risk:
$$\hat{f} = \arg\min_{f \in \mathcal{H}} \frac{1}{n}\sum_{i=1}^n \mathcal{L}(y_i, f(\mathbf{x}_i))$$
Key Properties
- Labeled data required
- Teacher signal guides learning
- Generalization to new examples
Challenge
Avoid overfitting to training data!
Python Example
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
model = LogisticRegression().fit(X_train, y_train)
print(f"Accuracy: {model.score(X_test, y_test):.2f}")Regression: Predicting Continuous Values
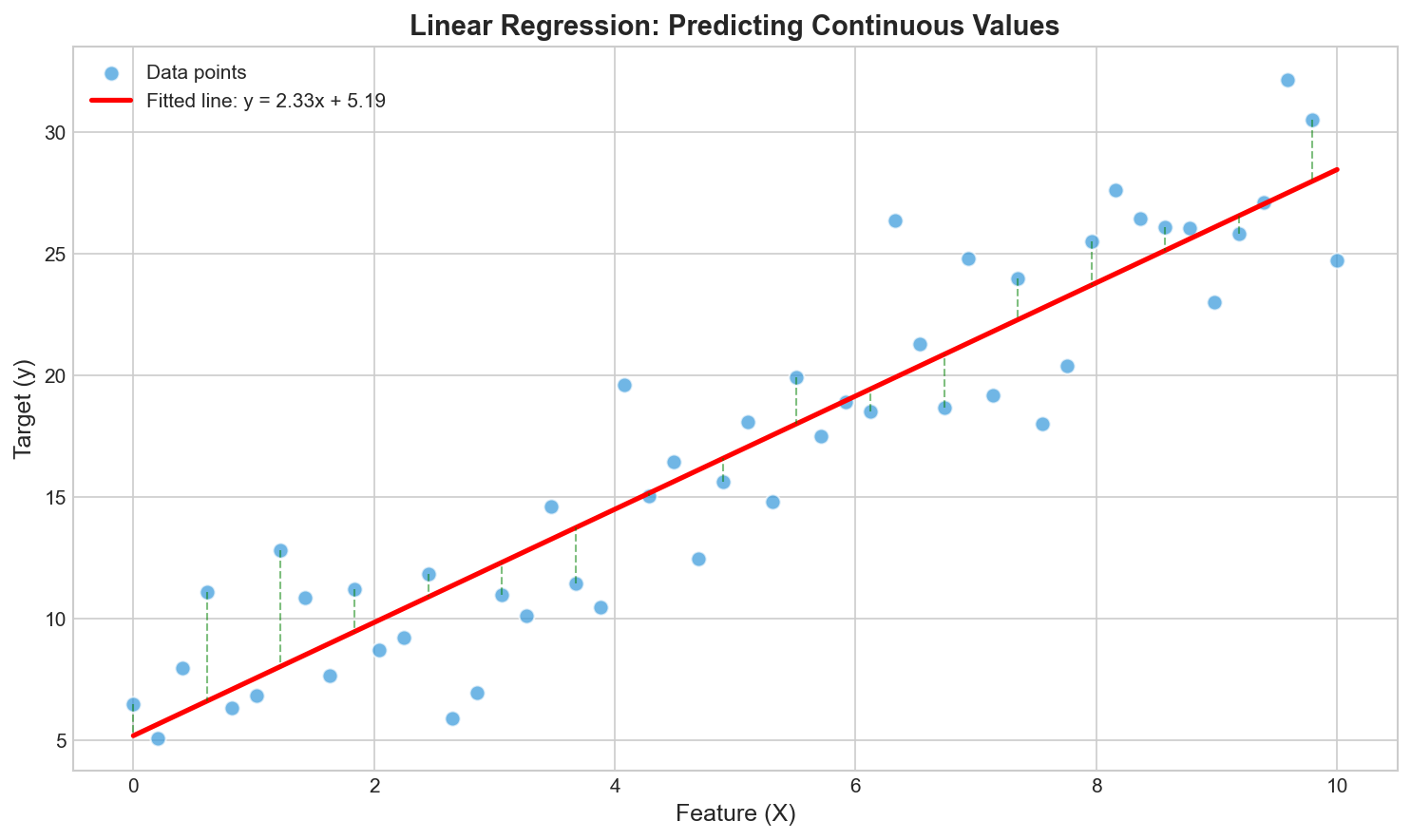
Input: $\mathbf{x} \in \mathbb{R}^d$
Output: $y \in \mathbb{R}$ (continuous)
Model: $\hat{y} = f(\mathbf{x}; \theta)$
Output: $y \in \mathbb{R}$ (continuous)
Model: $\hat{y} = f(\mathbf{x}; \theta)$
Loss Functions
- MSE: $\frac{1}{n}\sum (y_i - \hat{y}_i)^2$
- MAE: $\frac{1}{n}\sum |y_i - \hat{y}_i|$
Regression: Algorithms & Examples
Regression Algorithms
- Linear Regression
- Ridge/Lasso (regularized)
- Polynomial Regression
- SVR, Decision Trees
- Neural Networks
Real-World Examples
- House price prediction
- Stock forecasting
- Temperature prediction
- Sales forecasting
Python Example
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error, r2_score
model = LinearRegression().fit(X_train, y_train)
y_pred = model.predict(X_test)
print(f"MSE: {mean_squared_error(y_test, y_pred):.4f}")Classification: Predicting Categories
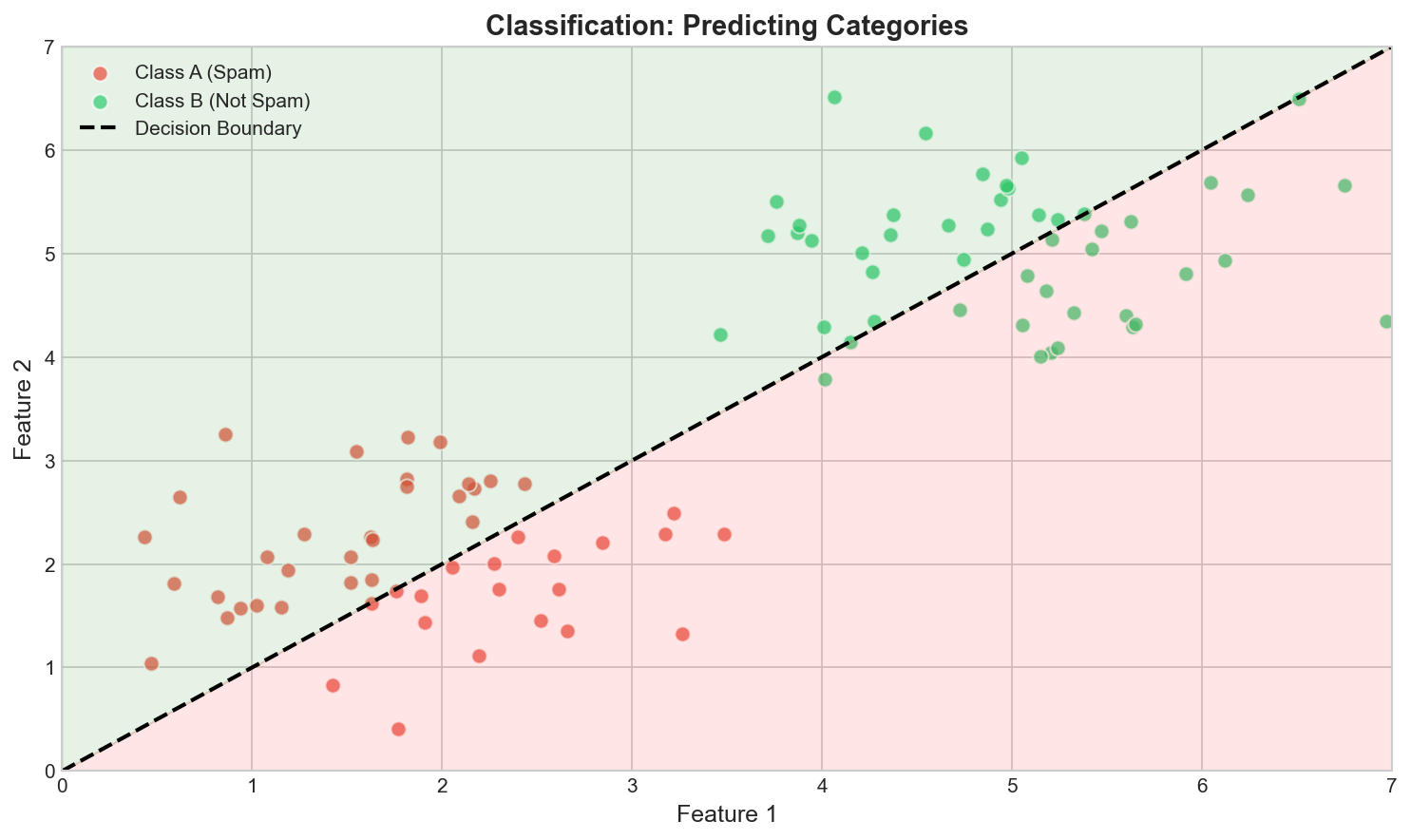
Input: $\mathbf{x} \in \mathbb{R}^d$
Output: $y \in \{1,...,K\}$ (discrete)
Model: $\hat{y} = \arg\max_k P(y=k|\mathbf{x})$
Output: $y \in \{1,...,K\}$ (discrete)
Model: $\hat{y} = \arg\max_k P(y=k|\mathbf{x})$
Types
- Binary: $K=2$ (spam/not spam)
- Multi-class: $K>2$ (digits 0-9)
- Multi-label: Multiple tags per item
Classification: Algorithms & Code
Classification Algorithms
- Logistic Regression
- Naive Bayes
- K-Nearest Neighbors
- Decision Trees
- Random Forests, SVM
Loss Functions
Cross-Entropy:
$$\mathcal{L} = -\frac{1}{n}\sum_{i} y_i \log \hat{y}_i$$Hinge Loss (SVM):
$$\mathcal{L} = \max(0, 1 - y\hat{y})$$Python Example
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import classification_report
clf = RandomForestClassifier(n_estimators=100).fit(X_train, y_train)
print(classification_report(y_test, clf.predict(X_test)))Unsupervised Learning
Definition
Learning from unlabeled data without explicit target outputs:- Input: Feature vectors $\{\mathbf{x}_1, …, \mathbf{x}_n\}$
- Output: None (discover structure)
Main Tasks
1. Clustering- Group similar data points
- Algorithms: K-Means, DBSCAN, Hierarchical
- Compress high-dimensional data
- Algorithms: PCA, t-SNE, UMAP
- Model the data distribution
- Algorithms: Gaussian Mixture Models
Key Characteristics
- No labels required
- Exploratory in nature
- Structure discovery
- Performance harder to measure
Applications
- Customer segmentation
- Anomaly detection
- Data visualization
- Feature extraction
- Compression
- Recommender systems
Challenge
How do we evaluate without labels?Supervised vs Unsupervised Comparison
| Aspect | Supervised | Unsupervised |
|---|---|---|
| Data | Labeled (X, y) | Unlabeled (X only) |
| Goal | Predict y from X | Find hidden patterns |
| Evaluation | Compare to true labels | Internal metrics |
| Examples | Classification, Regression | Clustering, PCA |
Clustering: Grouping Similar Data
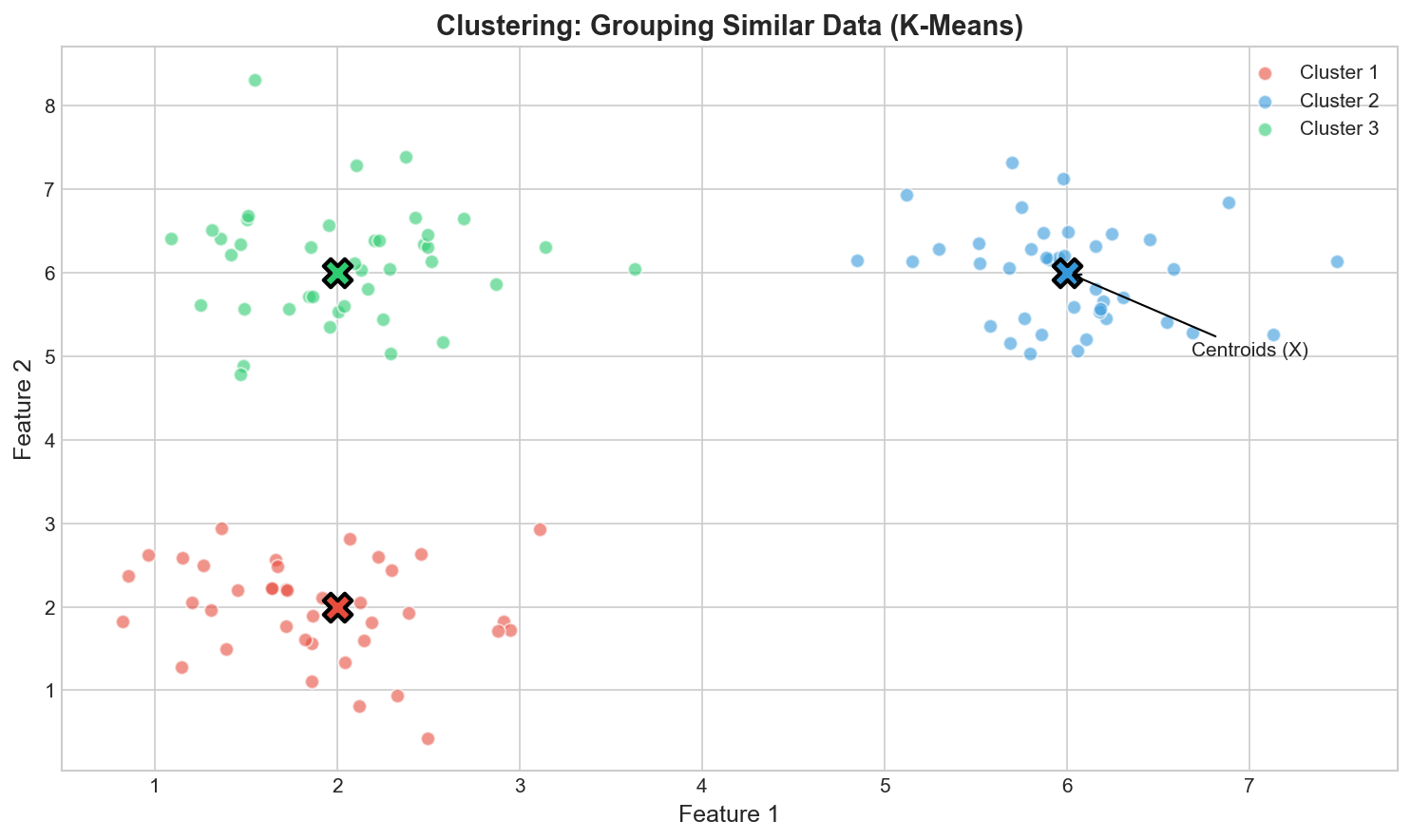
Goal: Group similar data points without labels
K-Means Objective
$$\min \sum_{i=1}^n \|\mathbf{x}_i - \mu_{c_i}\|^2$$- Initialize K centroids
- Assign points to nearest
- Update centroids
- Repeat until convergence
Clustering: Methods & Evaluation
Other Clustering Methods
- Hierarchical: Dendrogram
- DBSCAN: Density-based, finds arbitrary shapes
- GMM: Probabilistic, soft assignments
Evaluation Metrics
- Silhouette coefficient
- Davies-Bouldin index
- Calinski-Harabasz index
Python Example
from sklearn.cluster import KMeans
from sklearn.metrics import silhouette_score
kmeans = KMeans(n_clusters=3).fit(X)
score = silhouette_score(X, kmeans.labels_)
print(f"Silhouette: {score:.3f}")Dimensionality Reduction
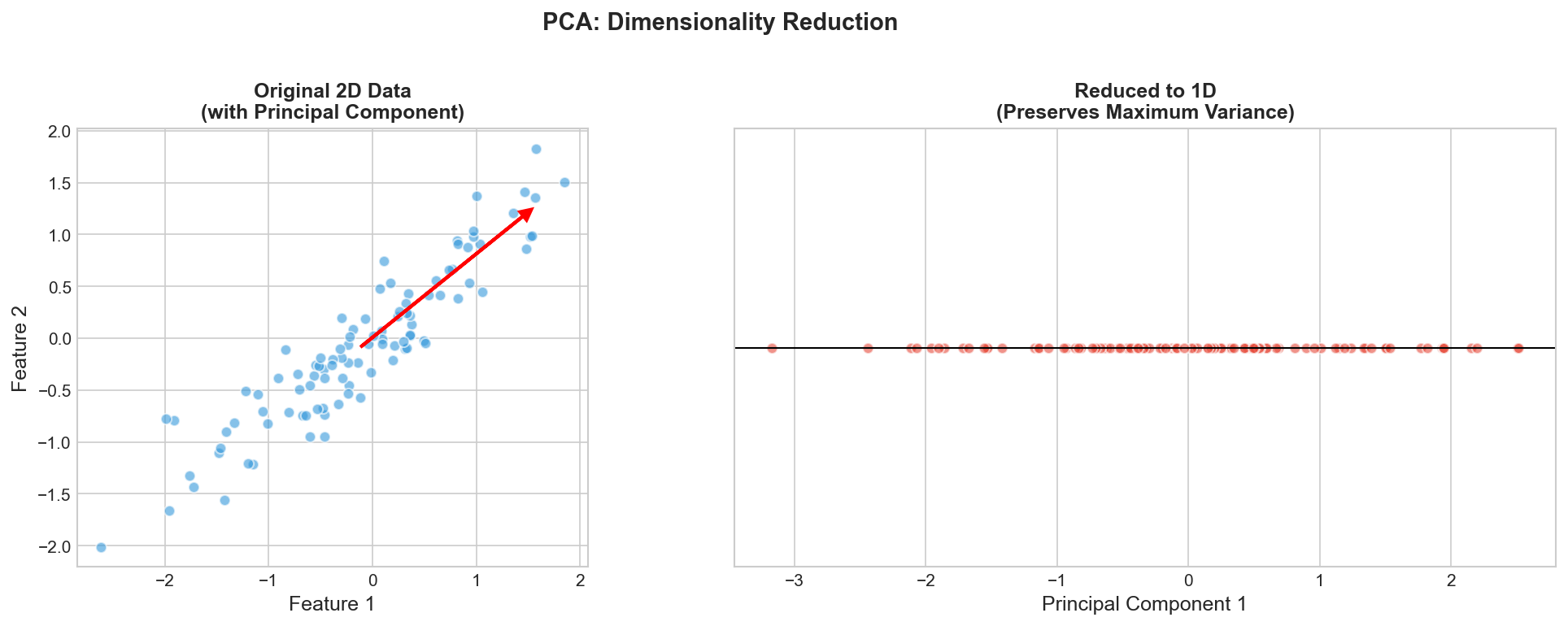
Goal: Compress high-dim data while preserving structure
Curse of Dimensionality
- Volume grows exponentially
- Data becomes sparse
- Overfitting risk increases
PCA & Other Techniques
PCA Algorithm
- Center data: $\tilde{\mathbf{x}}_i = \mathbf{x}_i - \bar{\mathbf{x}}$
- Compute covariance: $\mathbf{C} = \frac{1}{n}\mathbf{X}^T\mathbf{X}$
- Find eigenvectors of $\mathbf{C}$
- Project onto top $k$ eigenvectors
Other Techniques
- Linear: PCA, LDA, ICA
- Non-linear: t-SNE, UMAP
- Neural: Autoencoders
Python Example
from sklearn.decomposition import PCA
pca = PCA(n_components=2).fit(X)
X_reduced = pca.transform(X)
print(f"Variance explained: {pca.explained_variance_ratio_.sum():.2%}")Semi-Supervised Learning
Definition
Learning from both labeled and unlabeled data:- Labeled: $\mathcal{D}_L = \{(\mathbf{x}_1, y_1), …, (\mathbf{x}_l, y_l)\}$
- Unlabeled: $\mathcal{D}_U = \{\mathbf{x}_{l+1}, …, \mathbf{x}_{l+u}\}$
- Typically $l \ll u$ (few labels, many unlabeled)
Fundamental Assumptions
1. Smoothness Assumption- Nearby points share same label
- Data forms discrete clusters
- Points in same cluster have same label
- High-dim data lies on low-dim manifold
Common Approaches
Self-Training:- Train on labeled data
- Predict unlabeled data
- Add confident predictions to training set
- Iterate
- Multiple views of data
- Train separate classifiers
- Exchange confident predictions
- Construct similarity graph
- Propagate labels
Why Semi-Supervised?
Labels are expensive! (Human annotation, expert knowledge, time)Reinforcement Learning
"You can use a spoon to eat soup, but it's better to use a ladle. Learning is choosing the right tool." --- Yann LeCunDefinition
Learning through interaction with an environment:- Agent takes actions
- Environment provides states & rewards
- Goal: Maximize cumulative reward
Markov Decision Process (MDP)
Formal framework: $(\mathcal{S}, \mathcal{A}, P, R, \gamma)$- $\mathcal{S}$: State space
- $\mathcal{A}$: Action space
- $P(s'|s,a)$: Transition probabilities
- $R(s,a,s')$: Reward function
- $\gamma \in [0,1]$: Discount factor
RL vs Other Paradigms
Key Differences:- No direct supervision
- Delayed rewards
- Exploration vs exploitation
- Sequential decision making
- Trial and error learning
Classic Algorithms
- Q-Learning
- SARSA
- Policy Gradient
- Actor-Critic
- Deep Q-Networks (DQN)
- Proximal Policy Optimization (PPO)
Famous Applications
AlphaGo, robotics, game playing, autonomous drivingRL Example: Q-Learning
Q-Learning Algorithm
Goal: Learn optimal action-value function $$Q^*(s,a) = \max_\pi \mathbb{E}\left[\sum_{t=0}^\infty \gamma^t R_t \mid s_0=s, a_0=a, \pi\right]$$ Update Rule: $$Q(s,a) \leftarrow Q(s,a) + \alpha [r + \gamma \max_{a'} Q(s',a') - Q(s,a)]$$ where:- $\alpha$: Learning rate
- $r$: Immediate reward
- $s'$: Next state
- $\gamma$: Discount factor
Algorithm Pseudocode
- Initialize $Q(s,a)$ arbitrarily
- For{each episode}
- Initialize state $s$
- Repeat
- Choose action $a$ using $\epsilon$-greedy policy
- Take action $a$, observe $r, s'$
- $Q(s,a) \leftarrow Q(s,a) + \alpha [r + \gamma \max_{a'} Q(s',a') - Q(s,a)]$
- $s \leftarrow s'$
- Until{$s$ is terminal}
Key Concepts
Exploration vs Exploitation:- $\epsilon$-greedy: explore with probability $\epsilon$
- Balances trying new actions vs using known good ones
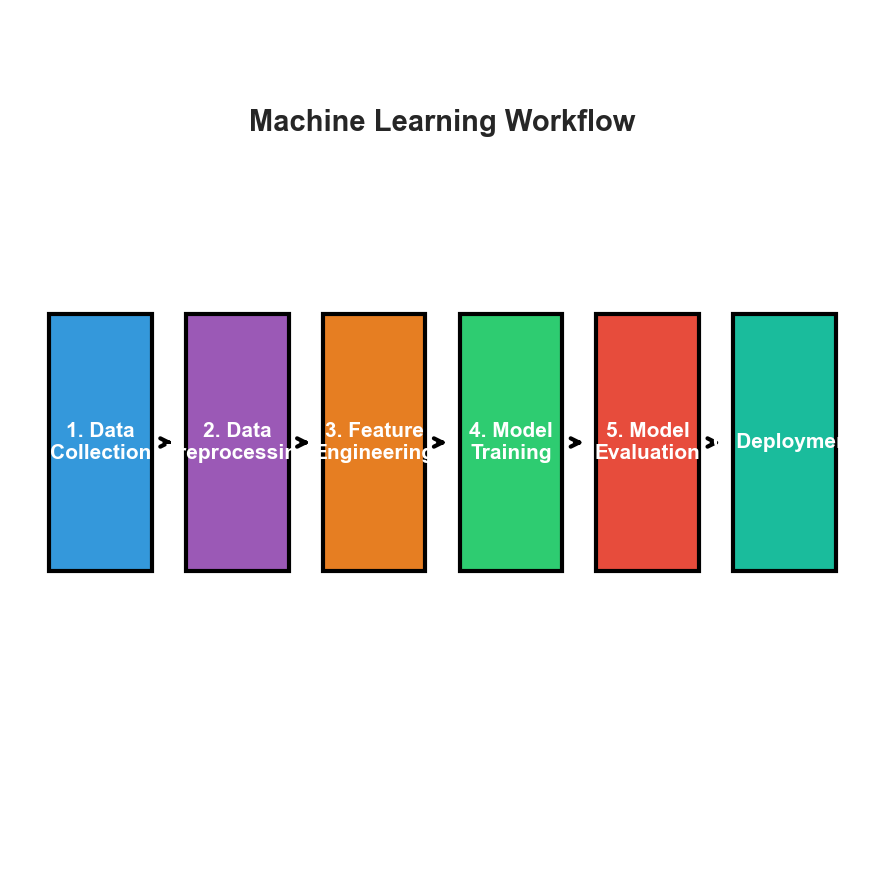
The ML Pipeline: From Data to Deployment
See visual diagram in lecture materials
Key Insight
ML is iterative! Model performance informs feature engineering, data collection, etc.Data Preprocessing: Cleaning & Scaling
Data Cleaning
- Missing values: Imputation or deletion
- Outliers: Detect and handle
- Duplicates: Remove
- Noise: Filter/smooth
Feature Scaling
Z-score: $z = \frac{x - \mu}{\sigma}$
Min-Max: $x' = \frac{x - \min}{\max - \min}$
Robust: Uses median/IQR
Why Scale?
Many algorithms (SVM, KNN, gradient descent) are sensitive to feature scales!
Feature Engineering & Train/Test Split
Feature Engineering
- Polynomial features: $x_1 x_2$, $x^2$
- One-hot encoding
- Date/time extraction
- Text vectorization (TF-IDF)
Train/Test Split
- Common: 80/20 or 70/30
- Cross-validation (k-fold)
- Time series: temporal split
Rule: Never train on test data!
Python Example
from sklearn.preprocessing import StandardScaler
from sklearn.impute import SimpleImputer
imputer = SimpleImputer(strategy='mean')
scaler = StandardScaler()
X_clean = scaler.fit_transform(imputer.fit_transform(X))Model Selection \& Training
Choosing a Model
Consider:- Problem type: Regression, classification, etc.
- Data size: Deep learning needs more data
- Interpretability: Linear models vs black boxes
- Training time: Real-time vs offline
- Prediction speed: Production requirements
No Free Lunch Theorem
Theorem: No single algorithm works best for all problems Implication: Must try multiple approaches and validate empiricallyStart Simple!
- Simple baseline (mean, majority class)
- Linear model
- More complex models
- Ensemble methods
Training Process
Optimization: Minimize loss function $$\theta^* = \arg\min_\theta \mathcal{L}(\theta; \mathcal{D})$$ Common Optimizers:- Gradient Descent
- Stochastic Gradient Descent (SGD)
- Adam (adaptive learning rate)
- RMSprop
Hyperparameter Tuning
Hyperparameters: Set before training- Learning rate, regularization strength
- Number of layers, hidden units
- Tree depth, number of trees
- Grid search
- Random search
- Bayesian optimization
Model Evaluation Metrics
Regression Metrics
Mean Squared Error (MSE): $$\text{MSE} = \frac{1}{n}\sum_{i=1}^n (y_i - \hat{y}_i)^2$$ Root MSE (RMSE): $$\text{RMSE} = \sqrt{\text{MSE}}$$ Mean Absolute Error (MAE): $$\text{MAE} = \frac{1}{n}\sum_{i=1}^n |y_i - \hat{y}_i|$$ R-squared (coefficient of determination): $$R^2 = 1 - \frac{\sum_i (y_i - \hat{y}_i)^2}{\sum_i (y_i - \bar{y})^2}$$ Range: $(-\infty, 1]$, closer to 1 is betterClassification Metrics
Accuracy: $$\text{Acc} = \frac{\text{correct predictions}}{\text{total predictions}}$$ Precision (positive predictive value): $$\text{Prec} = \frac{TP}{TP + FP}$$ Recall (sensitivity, true positive rate): $$\text{Rec} = \frac{TP}{TP + FN}$$ F1-Score (harmonic mean): $$F_1 = 2 \cdot \frac{\text{Prec} \cdot \text{Rec}}{\text{Prec} + \text{Rec}}$$ ROC-AUC: Area under ROC curveImportant
Choose metrics appropriate to your problem! Accuracy misleading for imbalanced data.When to Use Each Metric
| Metric | Use When | Avoid When |
|---|---|---|
| Accuracy | Balanced classes | Imbalanced data |
| Precision | False positives costly | Need recall |
| Recall | False negatives costly | Need precision |
| F1-Score | Balance precision/recall | Clear preference |
| RMSE | Penalize large errors | Robust to outliers |
| MAE | All errors equal | Large errors matter |
Bias-Variance Tradeoff
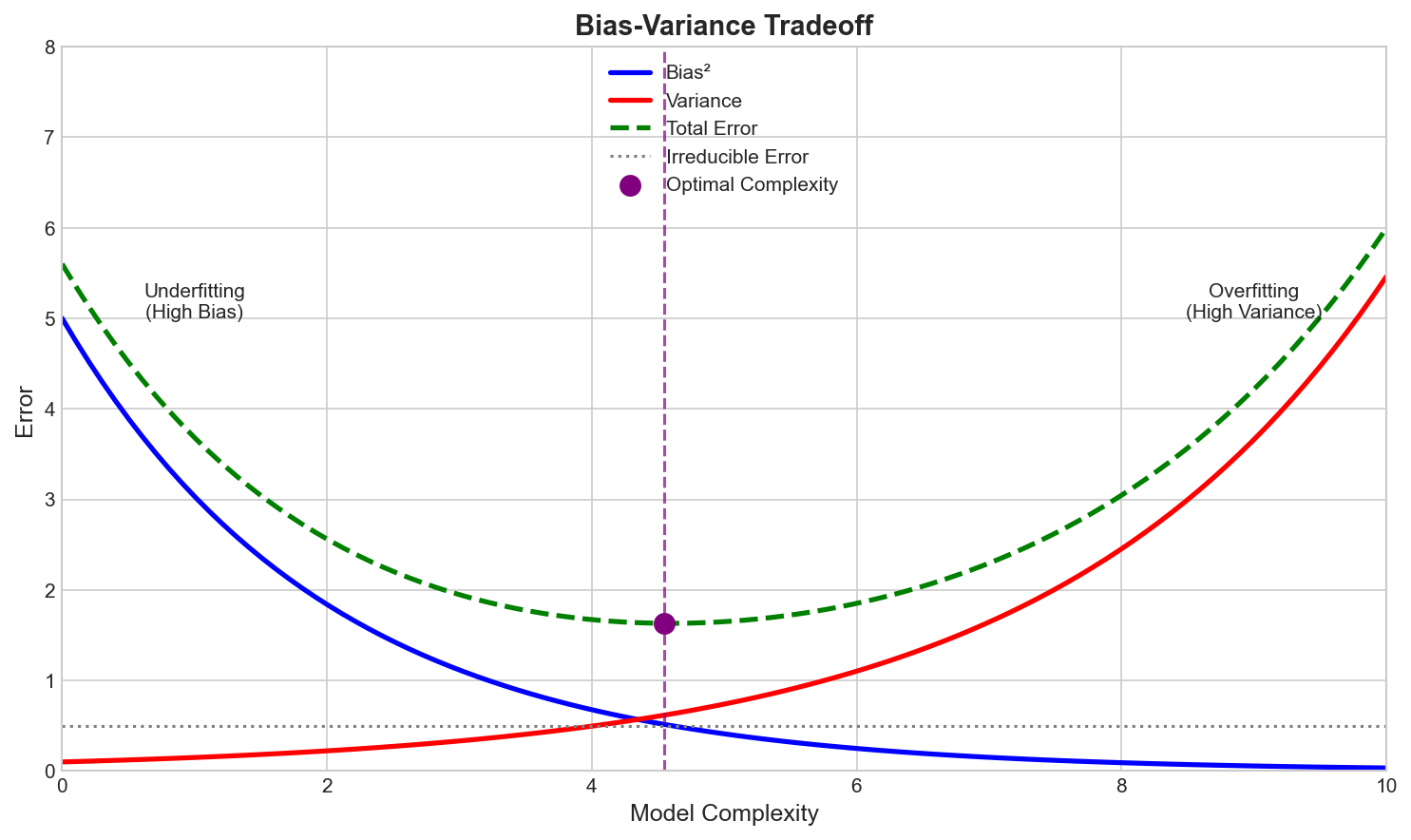
Error Decomposition
$$\text{Error} = \text{Bias}^2 + \text{Variance} + \text{Noise}$$- Bias: Error from wrong assumptions
- Variance: Sensitivity to training set
- Noise: Irreducible error
The Tradeoff
- Simple models: High bias, low variance
- Complex models: Low bias, high variance
Underfitting vs Overfitting
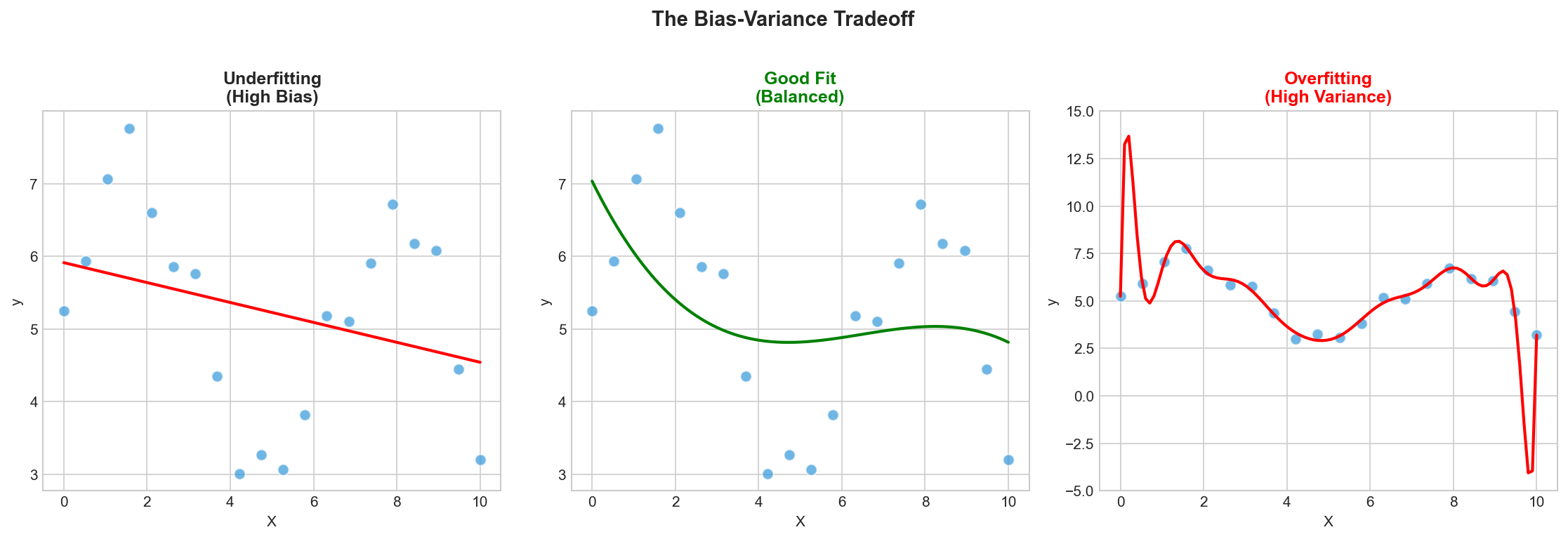
Underfitting
High train & test error
Fix:
- More features
- Complex model
- Less regularization
Overfitting
Low train, high test error
Fix:
- More data
- Regularization
- Simpler model
Regularization Techniques
L2 Regularization (Ridge)
Modified objective: $$\min_\theta \mathcal{L}(\theta) + \lambda \|\theta\|_2^2$$ where $\lambda > 0$ is regularization strength Effect:- Penalizes large weights
- Shrinks coefficients toward zero
- Improves generalization
- Handles multicollinearity
L1 Regularization (Lasso)
Modified objective: $$\min_\theta \mathcal{L}(\theta) + \lambda \|\theta\|_1$$ Effect:- Sparse solutions (some $\theta_i = 0$)
- Automatic feature selection
- More aggressive than L2
Elastic Net
Combines L1 and L2: $$\min_\theta \mathcal{L}(\theta) + \lambda_1 \|\theta\|_1 + \lambda_2 \|\theta\|_2^2$$ Benefits:- Sparsity from L1
- Stability from L2
- Best of both worlds
Python Example: Ridge and Lasso
from sklearn.linear_model import Ridge, Lasso, ElasticNet
# Ridge (L2 regularization)
ridge = Ridge(alpha=1.0)
ridge.fit(X_train, y_train)
# Lasso (L1 regularization)
lasso = Lasso(alpha=0.1)
lasso.fit(X_train, y_train)
# Elastic Net (L1 + L2)
elastic = ElasticNet(alpha=0.1, l1_ratio=0.5)
elastic.fit(X_train, y_train)
# Lasso creates sparse solutions (feature selection)
print(f"Non-zero coefficients: {sum(lasso.coef_ != 0)}")The Curse of Dimensionality
Problem Statement
As dimensionality $d$ increases:- Volume grows exponentially: $V \propto r^d$
- Data becomes sparse: Points far apart
- Distance metrics break down: All points equidistant
- Overfitting risk increases: More parameters to fit
Mathematical Insight
In high dimensions, volume concentrated in corners: $$\frac{V_{\text{corners}}}{V_{\text{total}}} = 1 - \left(1 - \frac{1}{2^d}\right)^{2^d} \approx 1 - e^{-1}$$ For unit hypercube, most volume is near edges!Data Requirements
To maintain density, need $n \propto c^d$ samples where $c > 1$Solutions
1. Dimensionality Reduction- PCA, t-SNE, UMAP
- Feature selection
- Filter methods (correlation)
- Wrapper methods (RFE)
- Embedded (Lasso, trees)
- L1/L2 penalties
- Early stopping
- Exponentially more needed
- Often impractical
Rule of Thumb
$n \geq 10 \cdot d$ for reliable modelsCMSC 173 Course Topics
Core Foundations
I. Overview (Today!)- Learning paradigms
- Applications
- Method of Moments
- Maximum Likelihood Estimation
- Linear Regression
- Lasso & Ridge
- Cubic Splines
- Bias-Variance Decomposition
- Cross-Validation
- Regularization
Advanced Methods
V. Classification- Logistic Regression, Naïve Bayes
- KNN, Decision Trees
- Support Vector Machines
- Kernel trick
- Principal Component Analysis
- Feedforward Networks
- CNNs, Transformers
- Generative Models
- K-Means, Hierarchical
- Gaussian Mixture Models
Learning Resources
Recommended Textbooks
Primary:- Murphy, K. P. (2022). Probabilistic Machine Learning: An Introduction. MIT Press.
- Bishop, C. M. (2006). Pattern Recognition and Machine Learning. Springer.
- Hastie et al. (2009). The Elements of Statistical Learning. Springer.
- Goodfellow et al. (2016). Deep Learning. MIT Press.
Online Resources
- Scikit-learn documentation
- PyTorch/TensorFlow tutorials
- Coursera ML courses (Andrew Ng)
- Stanford CS229 lecture notes
- ArXiv.org for research papers
Tools We'll Use
- Python 3.8+
- NumPy, Pandas, Matplotlib
- Scikit-learn
- Jupyter Notebooks
- PyTorch (for deep learning)
Installation
Ensure you have Python and required packages installed before next session!Best Practices in Machine Learning

Development Workflow
1. Start with baseline- Simple model first
- Establish minimum performance
- Change one thing at a time
- Track experiments
- Version control (Git)
- Cross-validation
- Hold-out test set
- Statistical significance
- Assumptions
- Hyperparameters
- Results
Common Pitfalls to Avoid
- Data leakage: Test data in training
- Ignoring class imbalance
- Not checking for overfitting
- Using wrong metrics
- Not scaling features
- Forgetting randomness: Set seeds!
- Over-engineering: Keep it simple
Reproducibility
Essential for science:- Set random seeds
- Document dependencies
- Share code & data (when possible)
- Report all hyperparameters
Ethics \& Responsible AI
"With great power comes great responsibility." --- Stan Lee (adapted from Voltaire)Ethical Considerations
Bias & Fairness:- Training data may contain biases
- Models can amplify discrimination
- Ensure fairness across groups
- Protect sensitive information
- Anonymization techniques
- Comply with regulations (GDPR)
- Explainable AI (XAI)
- Interpretable models
- Document limitations
- Adversarial robustness
- Prevent misuse
- Validate thoroughly
Societal Impact
Positive:- Healthcare improvements
- Scientific discoveries
- Accessibility tools
- Environmental monitoring
- Job displacement
- Deepfakes & misinformation
- Surveillance
- Autonomous weapons
Our Responsibility
As ML practitioners, we must:- Consider ethical implications
- Design inclusive systems
- Communicate limitations
- Prioritize societal benefit
Key Takeaways
What We Covered Today
- Definition of Machine Learning: Learning from data to improve performance
- Supervised Learning: Regression & classification with labeled data
- Unsupervised Learning: Clustering & dimensionality reduction
- Semi-Supervised Learning: Leveraging both labeled & unlabeled data
- Reinforcement Learning: Learning through interaction & rewards
- ML Pipeline: From data collection to deployment
- Key Challenges: Bias-variance tradeoff, overfitting, curse of dimensionality
- Best Practices: Systematic development, validation, ethics
Next Lecture
Parameter Estimation: Method of Moments & Maximum Likelihood EstimationPrepare for Next Session
Required Reading
Murphy (2022):- Chapter 4: Statistics (4.1-4.3)
- Chapter 5: Decision Theory (5.1-5.2)
- Chapter 1: Introduction (1.1-1.5)
- Chapter 2: Probability (2.1-2.3)
Practice Problems
- Review probability theory
- Linear algebra refresher
- Set up Python environment
- Install required packages
Questions to Ponder
- When would you choose supervised vs unsupervised learning?
- How do you decide on train/test split ratio?
- What metrics are appropriate for imbalanced datasets?
- How can we detect overfitting early?
- What are ethical concerns in your domain of interest?
Office Hours
Available for questions and discussion after class or by appointmentEnd of Module 00
Introduction to Machine Learning
Questions?